My first first-author publication was recently accepted into Marine Drugs, a peer-reviewed open access journal that broadly explores how marine natural products can be used as therapeutic agents, including for drug discovery and other biotechnology applications. In our study, we reported, for the first time, the venom-like gene profiles of four species of tube anemones and their potential as resources for novel therapeutic compounds. For this post I am going to present quite a bit of background beyond our study for context, so if you are interested in just a summary of the paper, scroll down until you see STUDY SUMMARY.
Big picture of the project: Explore new gene expression data from four species of tube anemones (cerianthids) for venom-like genes.
Basically, we were looking for venom genes in a pile of newly collected data from several understudied species of Cnidaria (phylum that includes jellyfish, sea anemones, corals, and other stingy, gelatinous critters). This study is the first (ever!) to look at the potential venoms that are in this group, expanding the knowledge of cnidarian venoms more broadly.
Before I get into the details, I need to shout out my co-authors of Team Cerianthid: Dr. Jason Macrander, Dr. Adam Reitzel, and Dr. Sérgio Stampar! Sérgio specifically works on all things tube-anemone (he has published over 20 papers on them!) and presented an amazing talk on the development and ecology of tube anemones at the dance party covertly disguised as Cnidofest, a conference specifically for the study of cnidarians. All three of these researchers are working with other cnidarian researchers to collect genetic data for tube anemones to learn more about their diversity, biology, ecology, and how they are related to other cnidarians. Dr. Stampar has discussed how the pelagic (swimming) larval stage of tube anemones are great models for understanding environmental effects. The “wow” factor from his talk was that the larvae from tube anemones (called cerinula larvae) look like jellyfish! Like, full on medusa with a bell and tentacles jellyfish!


Check out this amazing video by Dr. Stampar in one his studies on these incredible looking larvae!
Because I am interested in how venom changes from the benthic (attached to bottom) polyp to the pelagic (in the water column) medusae in jellyfish, cerianthids with a pelagic larva and a benthic polyp make an interesting and exciting comparative system! Unfortunately, this data does not (yet) exist, but Team Cerianthid continues to collect and sequence more cerianthid species for future research. I had some experience with looking for venom-genes in understudied cnidarians, i.e. my little hydroids, so I was asked if I would like to try looking for toxins in the four species they had data for. So began this really cool project that is now publicly available!
Let’s start with a bit of background on these animals. Tube-anemones are a group of cnidarians that belong in the same groups as corals and sea anemones (Class Anthozoa), though they superficially look like “typical” sea anemones. Cerianthids are in fact their own subclass of anthozoans, called Ceriantharia. Our understanding of the relationships between tube anemones and other anthozoans is still debated, i.e. tube-anemones might in fact be closely related to sea anemones or be what are called “basal” anthozoans, the earliest diverging group of anthozoans. Eventually I will write a post on the study of evolutionary relationships, called phylogenetics, and their importance within my work on venoms. For the purpose of this study, the evolutionary relationships between tube anemones and other anthozoans is fuzzy, which makes it difficult to know how the venoms of tube anemones and other sea anemones are related.
So why study their venoms anyway? We know they must have them right, they are cnidarians after all. Well, if that was the question we were asking, do they have venom, then yes, this would not be a very fruitful (or to be honest, interesting) study. But our questions were much more specific:
Initial question: What toxins can we identify within each of these four species?
Subsequent questions:
- Evolution/Biology: How similar or different are the toxins in each species to the others? What about to cnidarian venoms more broadly, especially other anthozoans?
- Function/Ecology: Can we use these toxin profiles to make educated guesses about how their venom works, both in terms of the mechanism of the toxin and the ecological role?
- Drug discovery: Are these toxins that could be used as novel drugs?
Now some nitty gritty: broadly, our study was a transcriptome-based exploration of venom-like genes within a species, which has been commonly done for many other neglected or small venomous species. So what is a transcriptome? One of the central tenets of biology is that DNA –> RNA –> protein, often called the central dogma. Now there are many more complex or nuanced pieces to this dogma, but generally it is presented as three levels of information: genome (DNA), transcriptome (RNA), proteome (proteins). DNA represents a genome-level study, which is looking at the genomes of multiple individuals of a species or single individuals more in depth to address various types of biological questions. While it is becoming more common to sequence the entire genome of a species, it is still expensive and very time consuming to make a good quality one. There is A LOT of data that can be studied from a genome, but for our study we do not necessarily need all the extra data. Now at any point certain parts of a genome are being expressed, and this is done when DNA is transcribed to RNA, specifically in our case messenger RNA (mRNA). These are little bits of sequence data that tell a researcher exactly what parts of the genome are being expressed at a particular time and in a specific place. The collection of this sequences data from RNA is called the transcriptome (i.e. the TRANSCRIBED bits of the genome). This is what we used for our study – the transcribed bits of the genome expressed in the tentacle tissue that was collected from four cerianthid species. Finally, studies at the protein level, the result of RNA being translated, is called proteomics. Often, transcriptome and proteome data are combined for a study of venom, especially for new species. But proteomics can be prohibitively expensive and often requires constant access to fresh animal tissue for sampling and troubleshooting, which is difficult when animals have to be constantly collected from the field. Transcriptomics also has problems, especially in terms of identifying venoms, but our hope is that this study is just the first of many on the venoms of these animals, and serves more as a platform that final report.
STUDY SUMMARY
I began my work directly form the assembled transcriptomes; my coauthors had already collected the animals (Ceriantheomorphe brasiliensis, Isarachnanthus nocturnus, Pachycerianthus borealis, and Pachycerianthus cf. maua) (check out Figure 1 in the paper for some fantastic pictures of these animals), collected the RNA from the tentacle tissues, and assembled the RNA sequences to make four transcriptomes. (For those new to reading scientific articles, there is often a section called “author contributions” at the bottom of the paper which describes what each co-author contributed to the study.) To use utilize the data in the transcriptome requires several bioinformatic techniques. Bioinformatics is the study of biological information using computational methods (i.e. various types of computer software/programs) to analyze biological questions. As mentioned above, our major question was to identify venom-like genes in each of these four species transcriptomes. I say venom-like genes because, without additional proteomics and/or other functional studies to determine how these toxins work and where they are located, we cannot confirm that these genes are in in fact true venom toxins. But I can use bioinformatics to narrow down good candidates for future studies.
One important aspect to mention is that the transcriptomes also have various quality control measures themselves, and there are many different types of programs, called assemblers, which take the short bits of sequenced RNA and put them together like puzzle pieces to make longer seuqences of DNA, called transcripts. I use transcripts to determine what might be a gene or not. Table 1 in our paper describes the number of small sequences we got for each species (reads), the number of transcripts or longer pieces the assembler could put together (transcripts), the number of genes that could be found in those transcripts (quality control measure to compare with other species), N50 value (quality control measure), and BUSCO, another quality control measure that tells you what percent of genes are found in your transcriptome that match to a conserved set of known genes in other animals. Based on these parameters, we knew all our transcriptomes were of good quality!
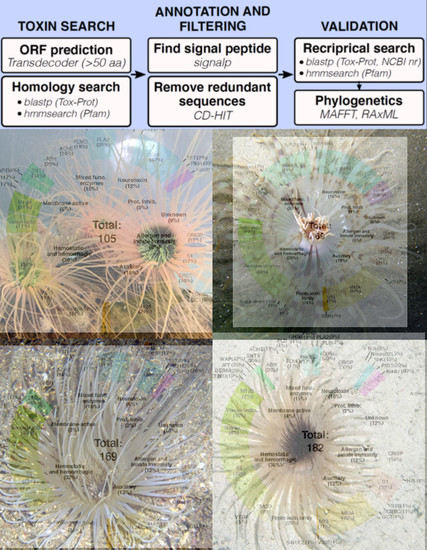
My customized pipeline (see top flow chart in the graphic abstract) used various pieces of software in three steps: create a pool of all possible genes within each of the four species, determine which genes could be toxins, and then sift through this set of potential toxins using various “quality-control” measures. I won’t get into the weeds with the exact tests and software I used, but one of the nice parts about open access manuscripts is all the methods are accessible for anyone to look at. Also, if anyone reading does have questions, please feel free to email me!
So, what can we do with this pool of venom-like genes? Well, we can do what every biologist learns to do very quickly during their career – count! First we counted the overall number of venom-like genes: around 100-200 proteins/peptides per species, which matches what we know for other cnidarian species. Weirdly, Isarachnanthus nocturnus had only 69 venom proteins, much lower than the 169, 182, and 105 from the other three species. Then we sorted all of the putative toxins from each species into seven different functional categories: neurotoxin, hemostatic and hemorrhagic toxins, membrane-active or pore-forming toxins, mixed function enzymes, protease inhibitors, allergen and innate immunity, and venom auxiliary proteins. Once that was finished, I calculated what the proportion of each of these “functional categories” is for each species, which provided a very rough estimate of how “functionally” similar the venom from each species was to the others. Unsurprisingly, like many cnidarian venoms, the majority of toxins belonged to hemostatic and hemorrhagic toxins, mixed enzymes, and auxiliary proteins (toxins that typically work synergistically with or assist in the production of venom proteins). What was VERY surprising is that neurotoxins, a common and often dominant component of other anthozoan venoms, is less than 20% of the venom profile in all four of the species of cerianthid. Two figures that help visualize these patterns are Figure 2 (series of venn diagrams) and Table 2 (listed number of specific toxin groups and proportions).
Our results section mainly encompassed going through each of the seven functional categories and referencing specific types of toxins we found within each of the four species. For instance, in the neurotoxins section, we talked about the various types of ShK domain containing toxins we found in each species. This included a figure (Figure 3) showing the actual sequence data, and the relevant parts that make up the ShK domain (like a puzzle piece that confers a specific function). These descriptions within the results are used to point out interesting protein families for future research, including potential candidates for future pharmacological testing or evolutionary oddities in terms of their presence in the venoms of tube anemones. For instance, within the pore-forming toxins section (toxins that literally make pores in cell membranes), three species of cerianthids had jellyfish (or CaTX/CrTX) toxins, which are highly potent pore-forming toxins typically found in box jellyfish. And yes, I mean those box jellies that cause really intense pain, possibly hospitalization or cardiac arrest form their stings. So what are these potent jellyfish toxins doing here? They have been found in other, less dangerous jellies, but not typically in anthozoans. Furthermore, we made a phylogenetic tree (Figure 4) of other known jellyfish toxins and the cerianthid jellyfish toxins. Phylogenetics, as I said, will have to be its own post, but the important thing about this tree is that the cerianthid toxins are not weird groups that fall on the outskirts of the tree, but are actually found within several other well-known toxin groups! That means these sequences are quite similar (i.e. related) and good candidates for further study. However, this is still not a guarantee that these toxins are present in the venom of these cerianthids as functional pore-forming toxins . I study this toxin family in my dissertation work, so hopefully in the near future I can talk more about them. There are also several more trees for other families of toxins that we found interesting, including actinoporins, stonutoxins, and phospholipase A2 – each of which could be of interest for future researchers studying cnidarian venoms given the placement of the cerianthid genes within the trees.
The discussion is where the results of the study are interpreted by the authors and put into context. Now, one of the major contributions is that we provided good quality data for four species of cerianthids, some of the first in this entire subclass, and this data can be used for a multitude of other experiments. Ours just happened to explore venoms, and overall, cerianthid venoms are WEIRD! In fact, based on the profiles that we found, if I had been asked if these toxins were from jellyfish or sea anemones, I would have said jellyfish! We tried to get as broad of a picture as possible of what toxins could be present, and even though this included many families of toxins common to anthozoans (i.e. actinoporins, sodium and potassium channel neurotoxins, small cysteine-rich peptides, etc.), we found very few. in any species. This is pretty interesting because, as I talked about above, we are not completely sure how cerianthids are related to other anthozoans. So in terms of our study, cerianthids might represent a more ancient venom profile for all anthozoans, or maybe they represent a loss of certain venom-like genes within anthozoans. Or maybe none of that is true! Until we have a better idea of the evolutionary history of this group, their placement, and a better understanding of their venom composition, we can only speak to broad patters that we have found from these results.
Moreover, we still do not know much about cnidarian venoms overall. In fact the vast majority of what we do know is from sea anemones. Because searching for toxins depends on previous knowledge from other toxin genes, it’s difficult to identify novel toxin genes, but perhaps more importantly it also biases the potential toxin genes we can find because we have a limited list to work with from the start. Remember, this is an entirely new group of animals we are exploring, and it will be important for future venom work to both continue trying to search for more venom-like gene in other species of cerianthid and other cnidarians more broadly.
All that said, I think what our study has done is actually quite wonderful – it opened up the door for many more questions to be asked! This not only includes what toxins are present or absent in cerianthid venoms, but how they work, how they are related to each other and other anthozoans, and what it tells us about cnidarian venom function and evolution more broadly. I am quite lucky to have been part of the first study on the venoms of these animals, and proud to have utilized some of the skills I have been building in graduate school. I am also grateful for the collaboration with #TeamCerianthid! If anyone has any questions about the study itself, the methods, or the experience taking on a project while in grad school, please feel free to reach out.
Paper: Klompen, A.M.L., Macrander, J., Reitzel, A.M., & Stampar, S.N. 2020. Transcriptomic Analysis of Four Cerianthid (Cnidaria, Ceriantharia) Venoms. Mar. Drugs, 18(8), 413. https://doi.org/10.3390/md18080413
Cover image: Pachycerianthus magnus from Stampar et al. 2019 (10.1038/s41598-019-42621-z)
References:
- Collins, A.G & Stampar S. N. 2014. No Bones Blog. https://nmnh.typepad.com/no_bones/2014/06/on-jellyfish-that-arent-jellyfish.html
- He, S., Grasis, J. A., Nicotra, M. L., Juliano, C. E., & Schnitzler, C. E. (2019). Cnidofest 2018: the future is bright for cnidarian research. https://doi.org/10.1186/s13227-019-0134-5
- Stampar, S. N., Broe, M. B., Macrander, J., Reitzel, A. M., Brugler, M. R., & Daly, M. (2019). Linear mitochondrial genome in Anthozoa (Cnidaria): a case study in Ceriantharia. Scientific reports, 9(1), 1-12. https://doi.org/10.1038/s41598-019-42621-z.

Fascinating! Three cheers for your hard work!
LikeLike
Thank you (:
LikeLike
WONDERFUL website! Thanks so much for this in-depth explanation of your research! I’m going to use it for my Invertebrate zoology class this (fall 2020) semester.
LikeLike
That is fantastic, thank you so much!! I am very happy to hear it will be useful for your class – best feedback I could hope for! My plan is to write about other projects I am working fairly soon. Good luck with your course this semester!
LikeLike